A groundbreaking study has mapped an elusive allosteric site in the KRAS protein, a key factor in many cancers. This discovery opens the door to the development of more effective and safer cancer treatments that target previously “untreatable” proteins. Credit: SciTechDaily.com
Researchers present for the first time a complete regulatory map of KRAS, offering a new blueprint for targeting “undruggable” proteins.
Researchers from the Center for Genome Regulation in Barcelona, Spain, and the Wellcome Sanger Institute near Cambridge, UK, have comprehensively identified allosteric control sites found in the protein KRAS. These are highly sought after drug targets and represent secret vulnerabilities that can be exploited to control the effects of one of the most important causes of cancer. The study presents the first complete regulatory map of any protein and was published Dec. 18 in the journal Science. Nature.
KRAS: An important target in cancer therapy
KRAS is one of the most frequently mutated genes in various types of cancer. It is seen in 1 in 10 cases of human cancer and is more prevalent in deadly types such as pancreatic and lung cancers. The protein is called the “Death Star” protein because it is globular and lacks a suitable site for drug targeting. For this reason, KRAS has historically been considered “untreatable” since his first discovery in 1982.
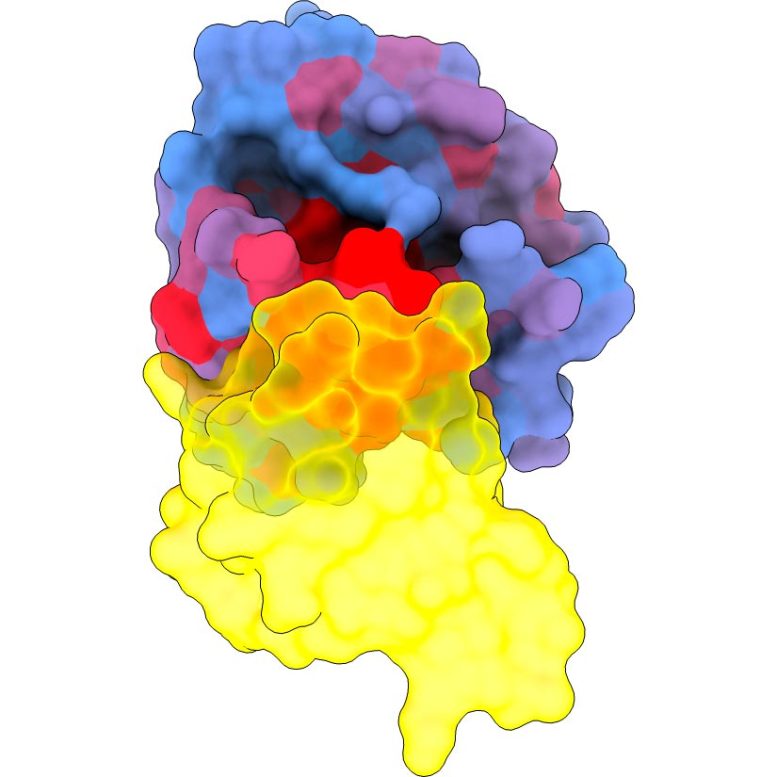
Three-dimensional image showing the human protein KRAS (blue) interacting with one of its major partners, RAF1 (yellow). The color gradient from blue to red indicates an increased potential for allosteric effects. Credit: Weng, Faure, Escobedo/Centro de Regulación Genómica
Allosteric sites: the key to controlling KRAS
The only effective strategy to control KRAS was to target its allostery communication system. These are molecular signals that work through the lock and key mechanism of your remote control. To control a protein, we need a key (a compound or drug) that opens the lock (active site). Proteins can also be influenced by secondary locks (allosteric sites) located elsewhere on their surface. When a molecule binds to an allosteric site, it causes a change in the protein’s shape, which can change the protein’s activity or ability to bind other molecules, including altering the internal structure of its main lock.
Challenges in allosteric drug development
Allosteric sites are often preferred for drug development because they provide higher specificity and reduce the potential for side effects. It also offers the possibility of more subtle changes in a protein’s activity and fine-tuning its function. Drugs that target the allosteric site are generally safer and more effective compared to drugs that target the active site.
However, allosteric sites are very difficult to understand. Despite 40 years of research, tens of thousands of scientific publications, and more than 300 published KRAS structures, only two of his drugs, sotorasib and adaglasib, have been approved for clinical use. is. Drugs act by binding to a pocket adjacent to the active site, inducing allosteric conformational changes that prevent protein activation.
Video showing the interaction between the human protein KRAS (blue) and one of its major partners, RAF1 (yellow), from different angles. The color gradient from blue to red indicates an increased potential for allosteric effects. This video focuses on novel and existing pockets that may control KRAS function through allosteric inhibition. Credit: Weng, Faure, Escobedo/Centro de Regulación Genómica
“It took decades to produce an effective drug against KRAS, in part because we lacked the tools to identify allosteric sites at scale, meaning we were blind to therapeutic target sites. That’s what you’ve been looking for. In this study, we demonstrate a new approach that allows you to systematically map allosteric sites across proteins. For drug discovery purposes, that means turning on a light and discovering the various ways a protein can be controlled. “It’s like figuring out how to do it,” explains Andre Fauré, Ph.D., staff scientist at the Center for Genome Regulation and co-author of the study.
Promising drug targets for KRAS protein
The authors of this study used a technique called deep mutation scanning to map allosteric sites. This involved creating more than 26,000 variations of the KRAS protein, changing just one or two components (amino acid)All at once. The research team looked at how these different KRAS mutations bind to six other proteins, including one that is important for KRAS to cause cancer. The researchers used AI software to analyze the data to detect allostery and locate known and novel therapeutic target sites.
“A unique selling point of our method is its scalability. In this study alone, we made over 22,000 biophysical measurements. A similar number to the sum done so far for all proteins before. DNA Sequencing and synthesis methodology. This is a significant acceleration and demonstrates the power and potential of this approach,” explains Chen-Chun Wen, lead author of the study and a postdoctoral fellow at the Center for Genome Regulation.
This technique revealed that KRAS has more potent allosteric sites than expected. Mutations at these sites disrupt the protein’s binding to all three major partners, suggesting that they can broadly inhibit KRAS activity. A subset of these sites is of particular interest because they are located in four distinct easily accessible pockets on the protein surface and represent promising targets for future drugs.
The study authors highlight “Pocket 3” as being particularly interesting. This pocket has received little attention from pharmaceutical companies until now because it is located far from the active site of KRAS.
The researchers also found that small changes in KRAS can dramatically change its behavior with partners, causing the proteins to prefer each other. This has important implications because it may lead to new strategies to control aberrant activity of KRAS without interfering with its normal function in non-cancerous tissues. Not using the regular version of his KRAS means a safer and more effective treatment with fewer side effects. Researchers can also use this knowledge to dig deeper into the biology of KRAS and explain how the protein behaves in different scenarios, which may determine its role in different types of cancer. It could be the key.
A new blueprint for drugging the ‘undruggable’
This study provides for the first time a complete map of allosteric sites in any protein. seed. This research shows that with the right tools and techniques, such as those used to map KRAS, new vulnerabilities in a variety of medically important proteins that have historically been considered “untreatable” can be discovered. It shows.
“The big challenge in medicine is not knowing which proteins cause disease, but rather not knowing how to control them. Our research aims to target these proteins and improve their activity. It represents a new strategy for accelerating the development of drugs that control allosteric sites. “This means that there is a high probability that Welcome Sanger Institute.
References: Chenchun Weng, Andre J. Faure, Albert Escobedo, and Ben Lehner, “Energetic and allosteric landscape of KRAS inhibition,” December 18, 2023. Nature.
DOI: 10.1038/s41586-023-06954-0