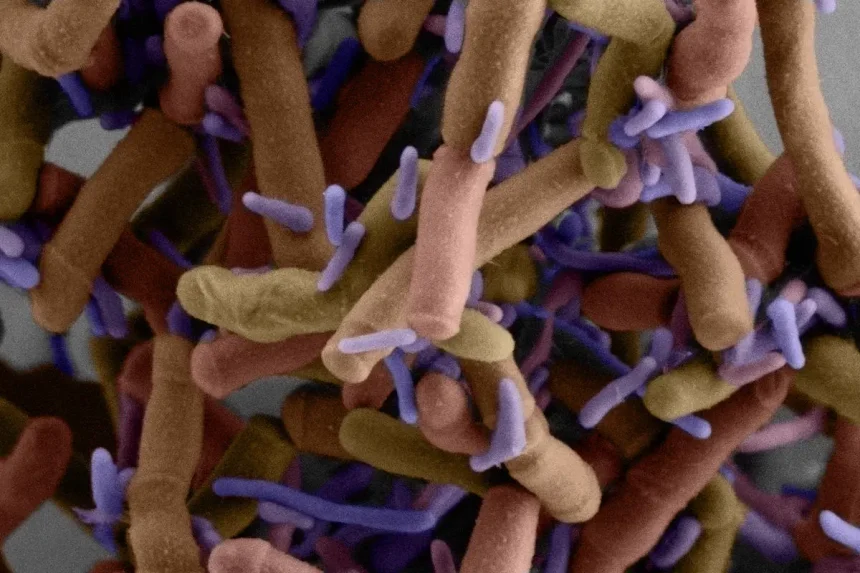
Scanning electron micrograph shows tiny purple Patescibacteria cells growing on the surface of much larger cells. A new study led by Joseph Moogs’ laboratory at the University of Wisconsin School of Medicine in Seattle has uncovered some of the molecular mechanisms thought to be behind their life cycle, genes, and their unusual lifestyle. Ta. These epibiotic bacteria are Southlakia epibionticum.Credit: Yaxi Wang, Wai Pang Chan, Scott Braswell/University of Washington
Scientists have discovered a gene essential to the unusual lifestyle of microscopic bacteria that live on the surface of larger bacteria.
Pathesibacteria are a mysterious group of micro-organisms with elusive survival methods. Scientists can only cultivate a handful of these types, but they are part of a diverse family found in many environments.
Several species of Patescibacteria that researchers can grow in the lab reside on the cell surface of another, larger host microorganism. Pathetic bacteria generally lack the genes necessary to make many of the molecules necessary for life. amino acid These include things that make up proteins, fatty acids that form membranes, and nucleotides. DNA. This has led researchers to speculate that many of them depend on other bacteria to grow.
In a recently published study, cell, researchers were able to glimpse for the first time the molecular mechanisms behind the pathecibacteria’s unusual lifestyle. This breakthrough was made possible by the discovery of how to genetically engineer these bacteria, an advancement that opens up a world of new research possibilities.
“Metagenomics can tell us which microbes live on and in our bodies, but DNA sequences alone are not particularly good at characterizing them,” said Nitin S. Barriga of the Institute. “It cannot provide insight into the beneficial or harmful activities of microbes that have never been detected.” He received an award in Systems Biology in Seattle and has contributed much computational and systems analysis to his work.

Epibiotic bacteria researcher Larry A. Gallagher sits behind a microscope in the microbiology lab at the University of Washington School of Medicine.Credit: S. Brook Peterson/University of Washington
“The ability to genetically perturb the puttyibacteria opens up the possibility of applying a powerful systems analysis lens to rapidly characterize the unique biology of obligate epibionts,” says another researcher for survival. Regarding organisms that must parasitize organisms, he added.
The team behind this research is led by Joseph Moogou’s laboratory at the National University’s Department of Microbiology. University of Washington The School of Medicine and the Howard Hughes Medical Institute were interested in pathetic bacteria for several reasons.
These are among the many poorly understood bacteria whose DNA sequence is seed-Rich microbial communities from environmental sources. This genetic material has been called “microbial dark matter” because little is known about the function it encodes.
microbial dark matter may contain information about biochemical pathways with potential biotechnology applications. cell paper. It also provides clues about the molecular activities that support microbial ecosystems and the cell biology of various microbial species that gather in those systems.
The Pathesibacteria group analyzed in this study belongs to the Saccharibacteria. They live in a variety of terrestrial and aquatic environments, but are best known for living in the human mouth. They have been part of the human oral microbiome since at least the Mesolithic and have been associated with human oral health.
In the human mouth, Saccharibacteria require a companion actinomycete to act as a host. To better understand the mechanisms that saccharibacteria employ to associate with their hosts, researchers used genetic engineering to identify all the genes essential for saccharibacterial growth.

Yaxi Wang, an epibiotic bacteria researcher at an anaerobic workstation in the microbiology lab at the University of Washington School of Medicine in Seattle.Credit: S. Brook Peterson/University of Washington
“We are very excited to have the first glimpse of the function of these rare genes in these bacteria,” said Moogs, a professor of microbiology. “By focusing future research on these genes, we hope to unravel the mystery of how saccharibacteria utilize host bacteria for growth.”
Potential host-interacting factors identified in this study include cell-surface structures that might help the Saccharibacterium to attach to host cells, and specialized proteins that might be used for nutrient transport. and secretory system.
Another application of the authors’ work was the generation of saccharibacterial cells expressing fluorescent proteins. The researchers used these cells to perform time-lapse microscopy fluorescence imaging of Saccharibacteria growing with host bacteria.
“Time-lapse imaging of cultures of Saccharibacteria and host cells reveals a surprising complexity in the life cycle of these rare bacteria,” said S. Brooke Peter, Senior Scientist in the Moogs Lab. Mr. Song said.
The researchers reported that some saccharibacteria act as mother cells by attaching to host cells and repeatedly budding to produce small swarms of offspring. These tiny cells start looking for new host cells. Some of the offspring became mother cells, while others appeared to interact non-productively with the host.
The researchers say that additional genetic engineering studies have opened the door to a broader understanding of the role of what they described as “a rich reservoir of microbial dark matter that these organisms contain,” yet to be imagined. We believe that it is possible to elucidate the biological mechanism that does not exist.
Reference: “Genetic manipulation of pathetic bacteria provides mechanistic insight into microbial dark matter and epibiotic lifestyle” Yaxi Wang, Larry A. Gallagher, Pia A. Andrade, Andi Liu, Ian By R. Humphreys, Serdar Turkarslan, Kevin J. Cutler, Mario L. Arrieta-Ortiz, Yachao Li, Matthew C. Raddy, Jeffrey S. McLean, Qian Kong, David Baker, Nitin S. Barriga, S. Brooke Peterson, Joseph D. Moogose, 7 Sept. 2023, cell.
DOI: 10.1016/j.cell.2023.08.017
This interdisciplinary and collaborative research was facilitated by a newly created research institute. Microbial Interaction and Microbiome Center (called its acronym mim_c), directed by Mougous. mim_c’s mission is to lower barriers to microbiome research and advance collaborative research through connections between like-minded researchers across disciplines. This led mim_c to join Moogs’ lab with oral microbiome expert Geoffrey MacLean from the Department of Periodontology at the University of Washington School of Dentistry.
The lead authors of this study were Yaxi Wang and Larry A. Gallagher of the University of Wisconsin Department of Microbiology. Senior authors were Baliga, Peterson, and Mougous. Biochemists Qian Cong and David Baker at the University of Texas Southwestern University and other researchers at the University of Wisconsin Institute of Medicine’s Institute of Protein Design also contributed to the study, along with McLean.
Moogs and Baker are researchers at the Howard Hughes Medical Institute. Moogs is the Lynn M. Garvey and Michael D. Garvey Endowed Chair at the University of Washington.
This study National Institutes of Healththe National Science Foundation, the Defense Threat Mitigation Agency, the Bill & Melinda Gates Foundation, and the Welch Foundation.